Program-Specific Researcher KATO, Yuki; Professor AKUTSU, Tatsuya and their research group “Developing an Extremely Fast Prediction Method for RNA-RNA Interaction” (Published in “Bioinformatics,” September 2010)
|
Program-Specific Researcher KATO, Yuki; Professor AKUTSU, Tatsuya and their research group (Laboratory of Biological Information Networks,Bioinformatics Center)
“Developing an Extremely Fast Prediction Method for RNA-RNA Interaction“
Published in “Bioinformatics,” September 2010 |

Dr KATO Y and Prof. AKUTSU T (from left) |
|
Yuki Kato1, Kengo Sato2, Michiaki Hamada3 4, Yoshihide Watanabe5, Kiyoshi Asai2 4 and Tatsuya Akutsu1 |
|
|
RNA (ribonucleic acid) is a stranded molecule that exists in living cells including humans. A few decades ago, RNAs were recognized that they have only passive roles such as messenger RNAs (mRNAs) that can be seen as the copy of information from DNA genes to produce proteins. |
|
| Recently, some of the non-coding RNAs (ncRNAs) have been pointed out that they have the mechanism of post-transcriptional regulation of gene expression, and considerable attention has been focused on the roles of ncRNAs to understand the mechanism of life. To take the regulatory mechanism as an example, the phenomenon that an ncRNA binds to its target mRNA so that the ncRNA inhibits/activates the synthesis of the corresponding protein has been observed (Figure 1). This phenomenon is called RNA-RNA interaction, and analysis of joint structure composed of two RNAs is important to elucidate the mechanism of RNA-RNA interaction. |
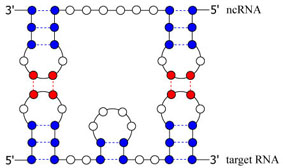 Figure 1. An example of RNA-RNA interaction. A pair of blue circles with a broken line represents an internal base pair, whereas a pair of red circles with a broken line indicates an external base pair.
|
|
A single stranded RNA is represented as a string consisting of four bases A, C, G and U, and its folding structure is constructed in such a way that the four bases selectively form hydrogen bonds. This mechanism of binding can be applied to the biding sites between two interacting RNAs. Several computational approaches have so far been proposed to RNA-RNA interaction prediction. Aim of these studies is to predict joint structures or binding sites given two RNA sequences. The framework of the existing methods, however, has a disadvantage such that their computational efficiency becomes worse when analyzing more complex joint structures. Therefore, efficient computational methods for predicting complex joint structures are still awaited. |
|
|
In this work, we develop RactIP, a fast and accurate prediction method for RNA-RNA interACTion using Integer Programming. RactIP solves the problem of maximizing expected accuracy of a predicted structure under an approximate probability distribution over a space of joint structures, using integer programming with threshold cut (Figure 2). |
|
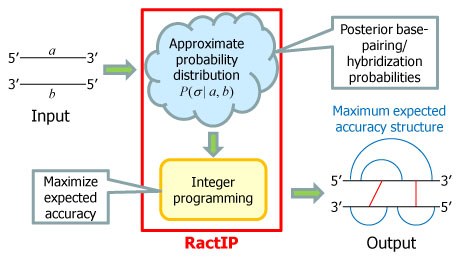 Figure 2. An overview of RactIP.
|
|
|
Experimental results on real interaction data show that RactIP is at least comparable in accuracy to several state-of-the-art methods. The important point to note is the drastic reduction of running time for prediction. For example, existing methods require 4000 seconds to predict some joint structures, whereas RactIP takes only 0.1 seconds for the same data, indicating that RactIP can run at most 40000 times faster than the earlier methods. The source code and binary code are freely available at http://www.ncrna.org/software/ractip/ |
|
|
This work has been accepted for oral presentation at ECCB2010, a major international conference on bioinformatics. Considering that the acceptance rate was less than 17%, we believe that our method will have a strong impact on the field of RNA bioinformatics. |
|
| This work was supported in part by Grant-in-Aid for Young Scientists (B) (KAKENHI) from Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan [#22700313 to Y.K., #22700305 to K.S.]. | |
|
Published in Bioinformatics, vol. 26, no. 18, pp. i460-i466, Sep. 2010. |
|
 Institute for Chemical Research, Kyoto University
Institute for Chemical Research, Kyoto University International Joint Usage Research Center
International Joint Usage Research Center